 
MetaboDirect traning session (December 1, 2021) You should arrive to a directory that contains several executables including Rgui, Rscript and Rcmd.Ĭlick on the address bar and copy the path to this directory. Select Rscript -> Open file location (see figure below). Search for Rscript in the search box of Windows. Locate the directory containing R executables Putting R in your system's PATH environmental variable (Windows)īy Christian Ayala, University of ArizonaĪ. It can be converted to an Apptainer container (formerly known as Singularity) so it can be run in the HPC. MetaboDirect's container is publicly available in DockerHub. 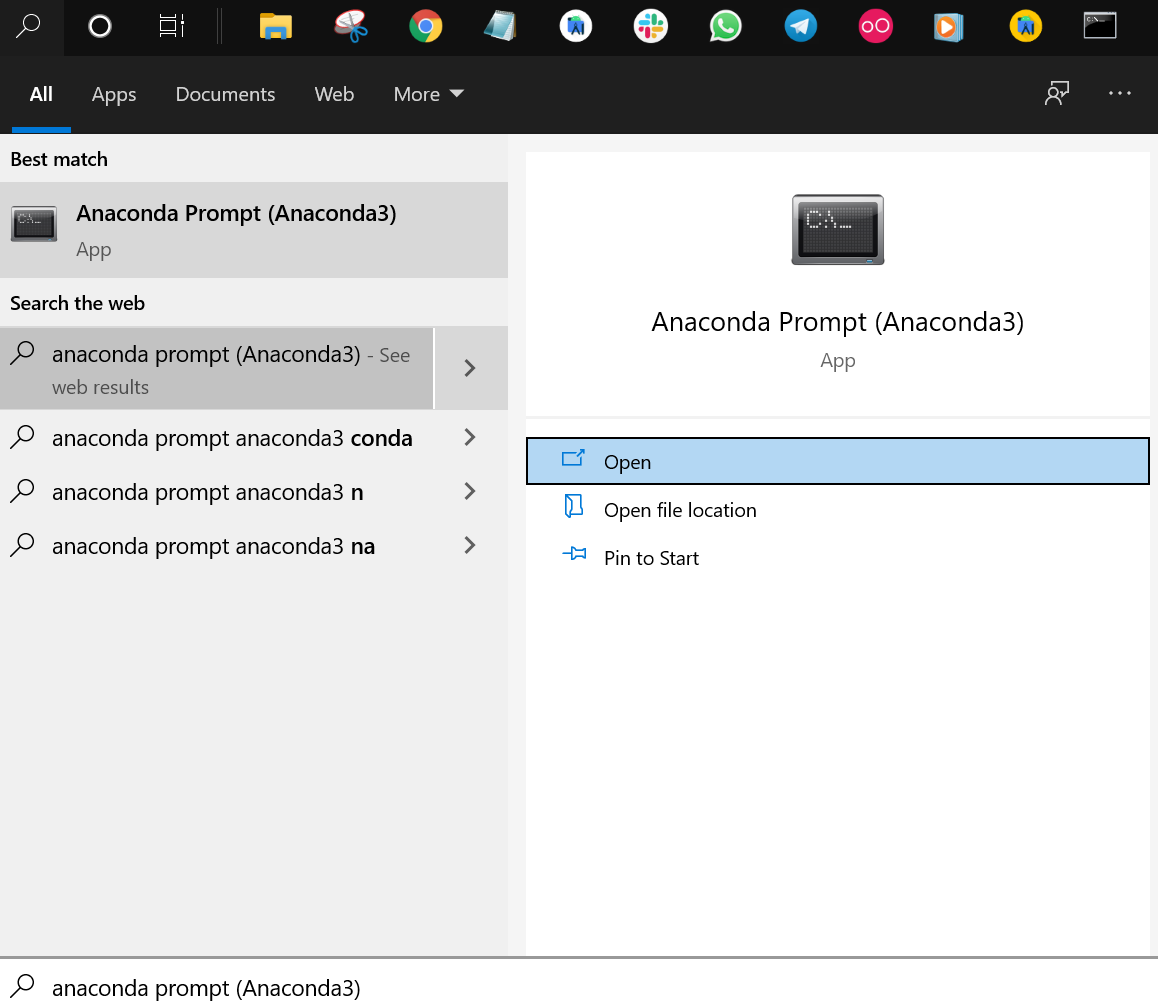
If you want to create an interactive session with more cpus or with more time you can check the help of the interactive command with interactive -help. 
TO start and interactive session use the following command: Running the container version of MetaboDirect in the UA HPCĬhristian Ayala-Ortiz Start an interactive sessionĪn interactive session is required to pull the container. > Thanks a lot for your suggestions Moira, I will definitely work on expanding the User Guide to provide a better description for the output files. I know you cite the paper, so anyone who wants to understand thoroughly can read it, but maybe you could give a 1-2 sentence definition of what it is/how to use it to get people started if they are completely new to the concept. You might also consider expanding a bit on the explanation of things like SPANS. For example, looking at the class and elemental composition files but I'm not sure what the units are for the numbers or how they were obtained, and in the error distribution file in diagnostics I'm not certain what the colum labeled "mean" is (I'm guessing "mean error" but it would be nice to have that spelled out somewhere). It would be great to have a bit more documentation on the automatically generated files in the help somewhere. If it keeps failing send me a screenshoot for me to see what is happening. You can also use `ls` to see what files are in your current directory. You can check which directory you are using `pwd` in the command line. Check that you are putting the names of the files correctly and that you are in the same directory where the files are. I'm unsure if I am writing the data/metadata names incorrectly or if there is a coding error. I keep on getting an error that says "metabodirect: error: the following arguments are required: DATA, METADATA" but I am putting the name of the files at the end of the metabodirect code. Always check that you have metabodirect last version (above is the command to udate it). > I imagine that you had to open RStudio tog et the plots because R was encountering and error (most likely because of the continuous values). Since the metadata is mostly comprised of categorical values (even time points can be considered factors) I think it should work well this way. I was having the same issue, so the new version of metabodirect will assume all the values in the metadata to be factors, so it does not hav any issues when plotting and coloring. To fix this I assign the columns to character values in r studio, but I wonder if there would be a way to specify in commandline whether data is continuous or categorical? And to adjust graphs accordingly?Īlso, the new updated version seems to be working on Mac without having to add python -m! Also, just a small thing, but if any of my metadata are numbers, it won't do the graphs because it can't plot continuous values. 
MetaboDirect requires at least one data file (`DATA`), one metadata file (`METADATA`) and at least one grouping variable to be defined with `-g` option.Īlso let me know any feedback or bugs that you find.įor the exploratory graphs, I always have to open Rstudio and run all the commands again in order to get all the graphs. Move to your Desktop using the Anaconda promptĭownload test data form the Github repository To install the latest version you can use Install the required packages specified in the **MetaboDirect** GitHub (). **(Mac Users)** Open your defaul terminal.`ls` and `wget` should already be installed by default **(For Windows Users)** Open the *Anaconda Prompt* in Administrator Mode, and enter the following commands: MetaboDirect user manual can be accesed ().ĭownload and install the Anaconda distribution of Python from (). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
